Post by JoseG on Mar 1st, 2016 at 2:01am
Hi,
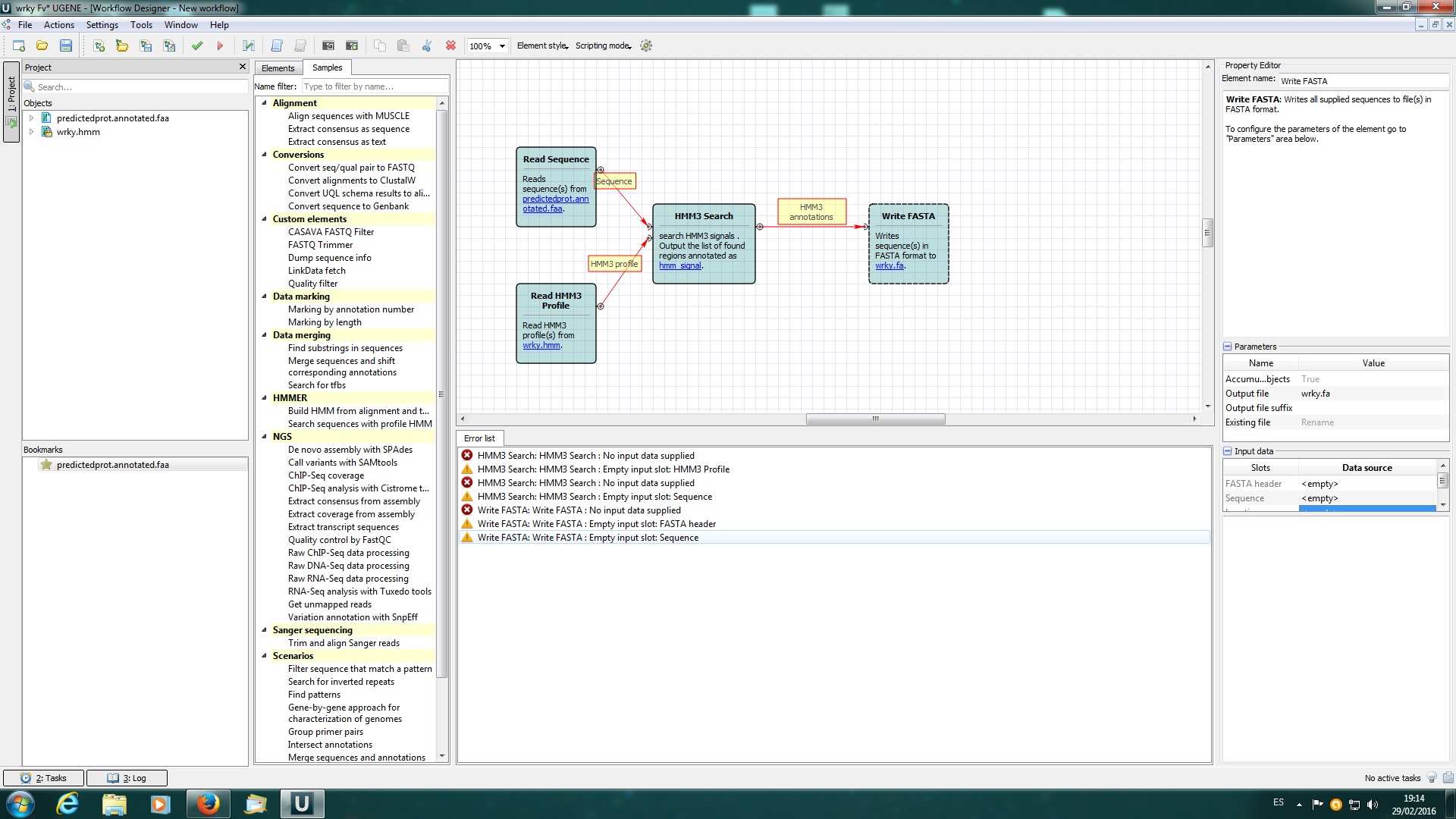
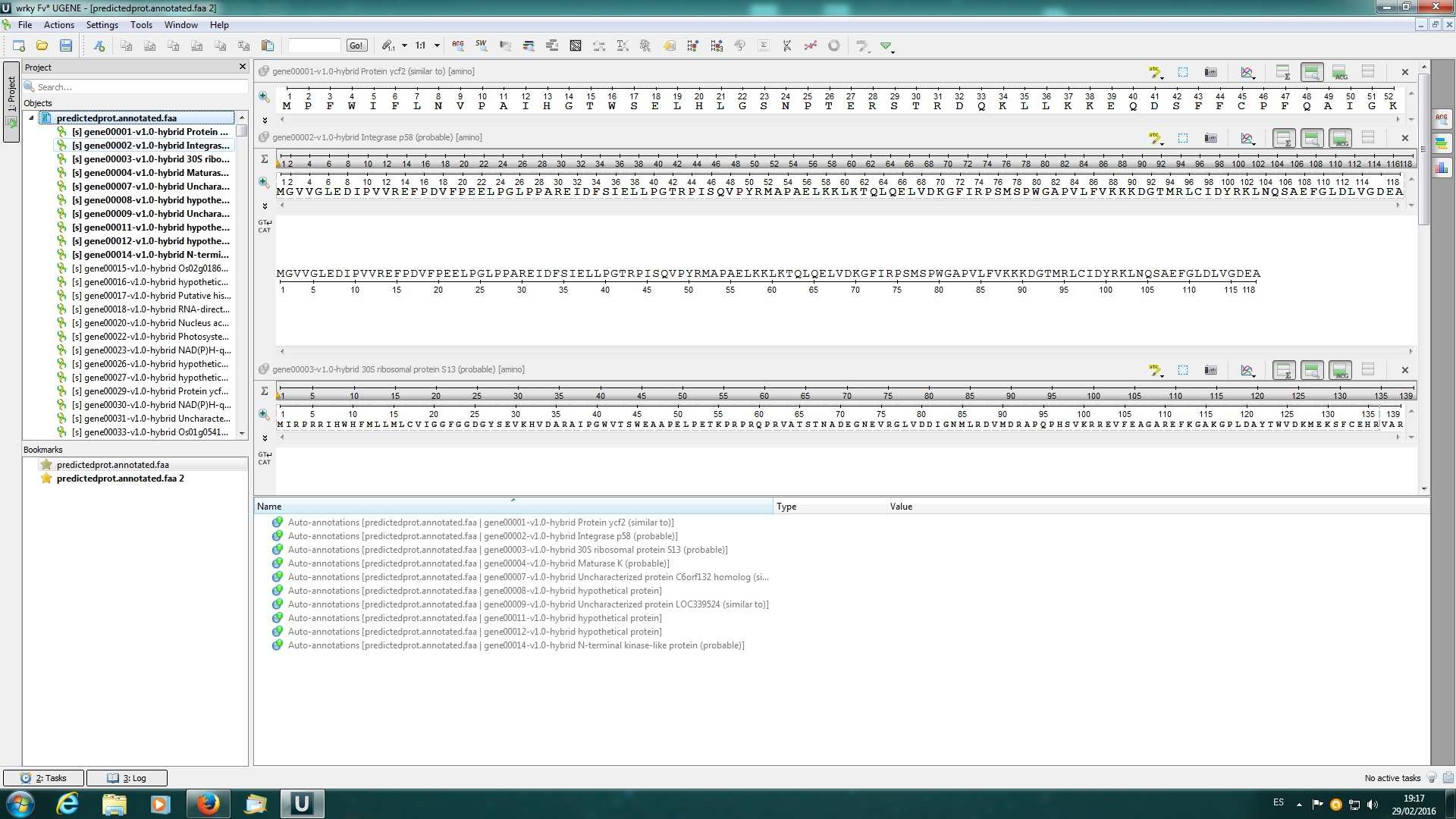
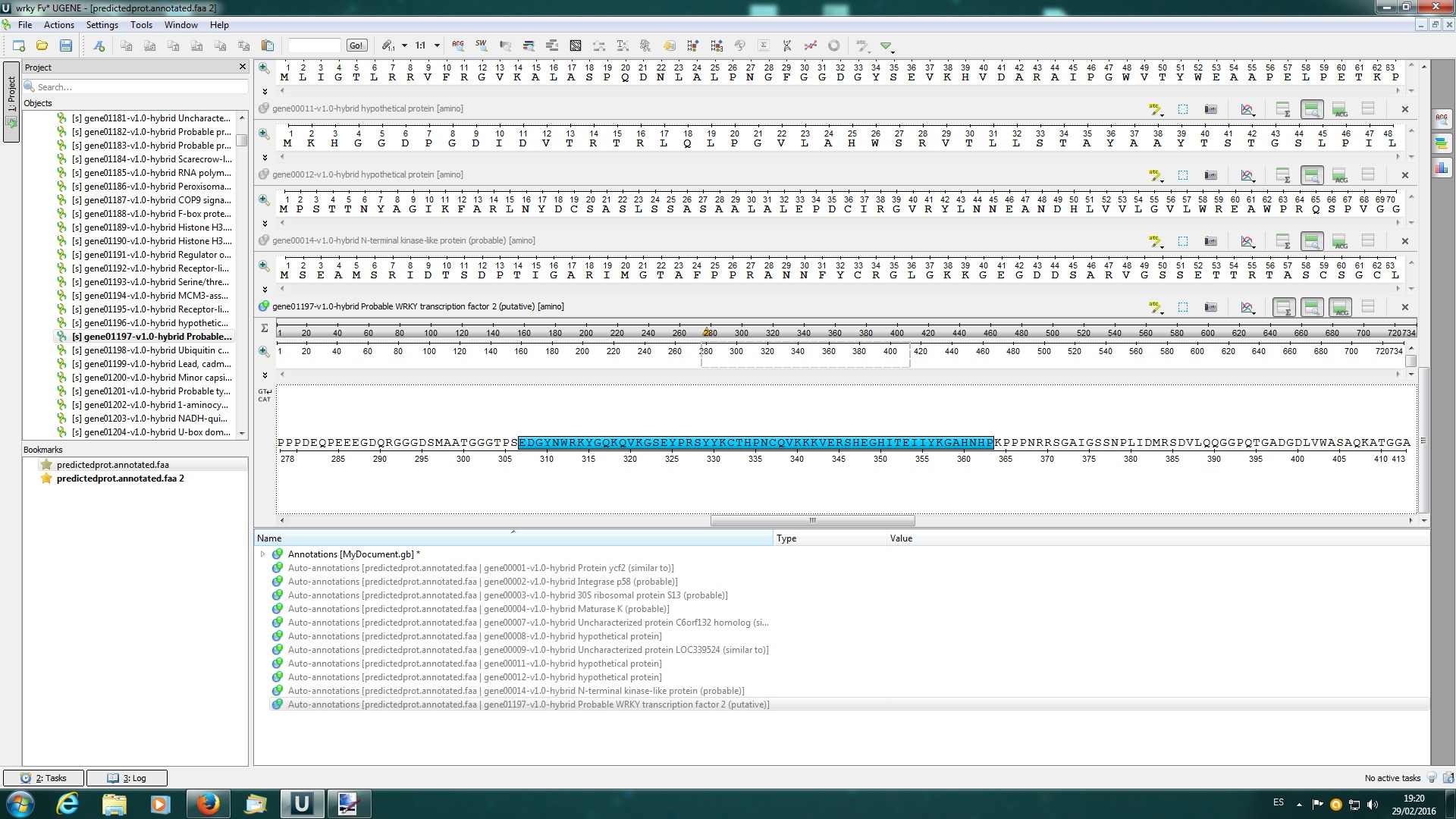
I attached 3 screenshots: first is the workflow (and errors) I build using the web documentation about "search sequences with profile hmm"; screenshots 1 and 2 are when I load the fasta file with more than 35000 seq (it only loads 14 in the viewer) and after I added manually one of them (positive) to perform the hmmer search with a pfam profile.
I cannot attach the fasta (very big) and hmmer profile files here but can provide download links if needed.
My question about filtering results is, supposing it can search for the whole list (screenshot1), how to get only the signal-positives? (if possible).
Any advice about how to do correctly the task?
Thank you
 ugeneworkflow.jpg (198 KB | 388
)
ugeneworkflow.jpg (198 KB | 388
)
 screenshot1.jpg (231 KB | 363
)
screenshot1.jpg (231 KB | 363
)
 screenshot2.jpg (242 KB | 369
)
screenshot2.jpg (242 KB | 369
)
I attached 3 screenshots: first is the workflow (and errors) I build using the web documentation about "search sequences with profile hmm"; screenshots 1 and 2 are when I load the fasta file with more than 35000 seq (it only loads 14 in the viewer) and after I added manually one of them (positive) to perform the hmmer search with a pfam profile.
I cannot attach the fasta (very big) and hmmer profile files here but can provide download links if needed.
My question about filtering results is, supposing it can search for the whole list (screenshot1), how to get only the signal-positives? (if possible).
Any advice about how to do correctly the task?
Thank you
 ugeneworkflow.jpg (198 KB | 388
)
ugeneworkflow.jpg (198 KB | 388
) screenshot1.jpg (231 KB | 363
)
screenshot1.jpg (231 KB | 363
) screenshot2.jpg (242 KB | 369
)
screenshot2.jpg (242 KB | 369
)
 https://forum.ugene.net/forum/YaBB.pl?action=downloadfile;file=hmm3.uwl (2 KB | 406
)
https://forum.ugene.net/forum/YaBB.pl?action=downloadfile;file=hmm3.uwl (2 KB | 406
)